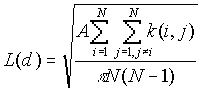 [1]
[1]K-Function
K-function is also called second-order analysis to indicate that the focus is on the variance, or second moment, of pairs of interevent distances. It considers all combinations of pairs of points. It compares the number of observed pairs with the expectation at all distances based on a random spatial distribution of points. The density of points, the borders, and the size of the sample are taken into consideration.
Input
Analysis
K-function analysis is a test of the hypothesis of CSR. The expected value of L(d) is d. The confidence interval in this analysis is generated by examining the specified number of permutations of randomly generated patterns of N points over the whole study area. If for any distance, the observed L(d) falls above or below the expected L(d) the null hypothesis of CSR can be rejected at an appropriate level of significance. The level of significance is determined by the confidence envelope. An observed L(d) below the envelope indicates that the points are dispersed at that distance, whereas an observed above the envelope indicates that clustering is present at that distance.
Formula
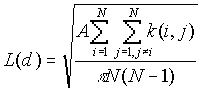 [1]
[1]
where:
A is the study area,
N is the number of points
d is the distance
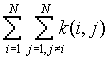 is the number of j points within distance d of all i points
is the number of j points within distance d of all i points
![]() is the weight, which is estimated by
is the weight, which is estimated by
a) If no edge corrections,
![]() in case d(i,j) £ d
in case d(i,j) £ d
![]() otherwise
otherwise
b) If a point i is closer to one boundary than it is to point j, the border correction is employed
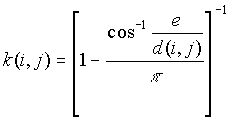 [2]
[2]
where e is the distance to the nearest edge.
c) If a point i is closer to two right angle boundaries than it is to point j, the weighting formula is
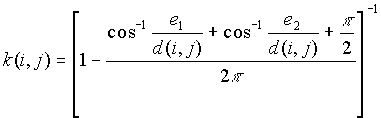 [3]
[3]
where
Output
The output table shows the distance, the observed
L(d), the minimum envelope L(d), and the maximum envelope L(d).|
Distance |
Observed L(d) |
Minimum L(d) |
Maximum L(d) |
|
: : : |
Limitation
The boundary correction formulas used here are inappropriate for irregular borders. In this program we assume the study area is a rectangle or square.
Example
For this example we will consider the distribution of the county seats of New Mexico. A map of New Mexico County Seats is shown in Figure 1.
Figure 1: New Mexico County Seats

There are a total of 32 points in this analysis. The input file is arranged in 32 rows of X, Y coordinates, and W values. Recall, all of the W values are equal to 1. A portion of the input data file is shown in Table 1. New Mexico is approximately 500km per side, so we will set our maximum study distance at 250km. We choose 25 increments so that we will calculate the observed L(d) and confidence interval for every 10km. 99 permutations are used for creating the confidence envelope in order to test the null hypothesis at the a=0.01 level.
Table 1: Input File
X Y W 219 338 1 27 189 1 418 156 1 408 538 1 534 262 1 438 272 1 . . . . . . . . . 73 83 1
A graph of the K-function results is shown in Figure 2, and the output file is given in Table 2. The observed L(d) is 0 for 10km and 20km because the closest pair of points is approximately 29km apart. At a distance of 30km, the observed L(d) falls within the generated confidence interval. However, for distances between 40km and 90km the observed L(d) lies outside of the confidence interval. This indicates that we can reject the null hypothesis of CSR. Also, since the observed L(d) is less than the Minimum L(d), this implies that we have a statistically significant dispersed or regular distribution of points.
Figure 2: Graph of K-Function Results
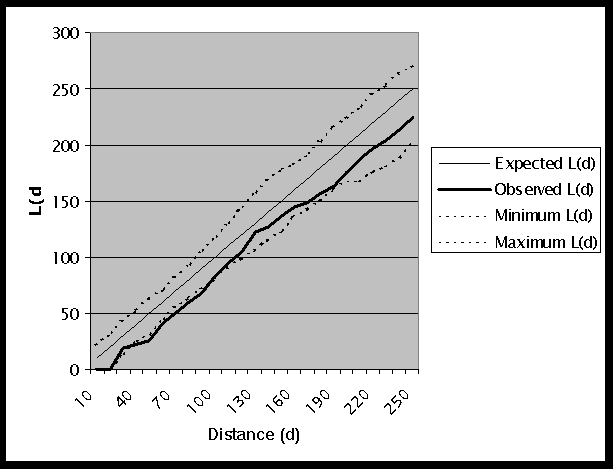
Table 2: Output File
The input data file: nmlung.txt The total number of points: 32 The minimum x coordinate: 27.000000 The maximum x coordinate: 534.000000 The minimum y coordinate: 30.000000 The maximum y coordinate: 538.000000 The total area: 257556.000000 The maximum search distance: 250.000000 The step size: 10.000000 The number of permutation for significance envelope:99 Distance Observed L(d) Minimum L(d) Maximum L(d) 10.00000 0.00000 0.00000 22.26797 20.00000 0.00000 0.00000 32.45112 30.00000 18.71896 12.85642 43.69063 40.00000 22.70875 26.09123 53.86896 50.00000 26.09549 31.49167 62.89654 60.00000 41.24103 44.67619 70.69446 70.00000 50.27186 55.09439 82.33472 80.00000 59.81204 64.09466 93.76176 90.00000 68.19498 72.26383 105.88598 100.00000 83.30301 81.39759 117.37930 110.00000 94.72131 90.86065 129.75777 120.00000 104.07702 98.57581 144.20721 130.00000 122.02663 106.60428 157.24152 140.00000 126.68748 115.87605 168.35863 150.00000 136.43529 123.72436 178.46855 160.00000 144.70865 136.44028 183.43776 170.00000 149.38940 143.72928 191.86183 180.00000 156.48955 150.63394 203.29807 190.00000 163.13176 160.98737 216.70522 200.00000 175.23454 167.31747 224.97855 210.00000 187.79141 168.27689 234.22479 220.00000 196.62003 176.33281 245.17390 230.00000 204.51863 180.89360 254.47212 240.00000 212.96201 189.13161 264.24158 250.00000 224.88269 201.79834 271.39911
References
Boots, Barry N. and Getis, Arthur, 1988, Point Pattern Analysis, Sage University
Paper series on Quantitative Applications in the Social Sciences, series no.07-001, Beverly Hills: Sage Publications
National Cancer Institute Biometry Research Group Datasets
http://dcp.nci.nih.gov/bb/datasets.html