![]() [1]
[1]
where N is the total number of points
Refined Nearest Neighbor Analysis
Refined nearest neighbor analysis involves comparing the complete distribution function of the observed nearest neighbor distances, F(di £ r), with the distribution function of the expected nearest neighbor distances for complete spatial randomness (CSR), P(di £ r). CSR is generated by means of two assumptions: 1) that all places are equally likely to be the recipient of a case (event) and 2) all cases are located independent of one another. This procedure differs from Nearest Neighbor Analysis in that the basis for border correction is taken from literature that recommends that a "guard area" represent an objective view of border conditions. (see Boots and Getis 1988)
Input
You’ll be asked to enter the input data file, which should contain N rows of X, Y coordinates, and W values. Make all W values equal to 1, representing points.
Analysis
Refined nearest neighbor analysis is used to test the null hypothesis of CSR. If for each distance, F(di £ r) > P(di £ r), a clustered pattern is indicated, whereas P(di £ r) < P(di £ r) indicates a regular pattern. The largest absolute distance (dr) between the F(di £ r) and the P(di £ r) is found. The significance of dr is then measured using a Monte Carlo test. If for this significant value, dr, F(di £ r) > P(di £ r) clustering is implied.
Formula
![]() [1]
[1]
where N is the total number of points
![]() [2]
[2]
Where:
e is the constant 2.178283
p is the constant 3.141593
r is the specified distance
l is the estimated point density (N/A)
c) dr
![]() [3]
[3]
where max | | means the largest absolute value obtained for corresponding values of r
Output
The output file includes three parts: the first part lists a) the input data file, b) the total number of points, c) the minimum and maximum of X, and Y coordinates, and d) the size of study area. The second part is a table in the following form:
|
r (distance) |
n1 (observed number of points for which di £ r) |
n2 (observed number of points for which ui < r < di) |
F(r) (observed proportion P(di £ r)) |
P(r) (expected proportion P(di £ r)) |
|F(r) - P(r)| (absolute value of the difference) |
|
: : : |
Limitation
In this program, the study area is a regular rectangle or square. Area (A) is calculated by (Xmax – Xmin) * (Ymax – Ymin).
Example
For this example of Refined Nearest Neighbor Analysis we will consider Figure 1, which shows incidents of larynx cancer the Chorley and South Ribble area of Lancashire, England. This data is available on Peter Diggle’s web site (http://www.maths.lancs.ac.uk/~diggle/pointpatterns/Datasets/). The null hypothesis (H0) is that the points are in a CSR pattern. The area used in the analysis is shown by the rectangle in Figure 1. The Xmin, Ymin, Xmax, and Ymax are shown in the output file (Table 2).
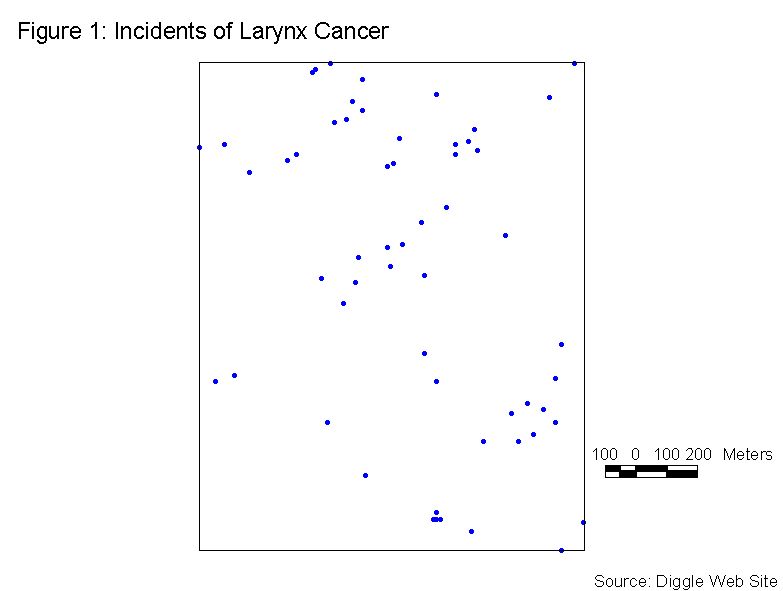
There are a total of 58 points in this sample. The input file is arranged in 58 rows of X, Y coordinates, and W values. For Refined Nearest Neighbor Analysis, all of the W values are set to 1. A portion of the input data file is shown in Table 1.
Table 1: Sample of input data file
X Y Z 35320 42800 1 35310 42230 1 34910 41850 1 35260 42080 1 35300 42150 1 35230 42660 1 36000 42850 1 34960 42500 1 35690 42570 1 . . . . . . . . . 34850 41830 1
For these data, table 2 shows that dr is 0.371. The results of the Monte Carlo test, a simulation run 99 times, indicates that this is significant at the 0.01 level. The null hypothesis of CSR can be rejected for a =0.01. This dr occurs at a distance of 63.246 meters, where the F(di £ r) > P(di £ r). This implies that the pattern is clustered.
Table 2: Output File
The input data file: d:\ppa\lardat.txt
The total number of points: 58
The minimum x coordinate: 34800.000000
The maximum x coordinate: 36030.000000
The minimum y coordinate: 41290.000000
The maximum y coordinate: 42850.000000
The total area: 1918800.0000
r, n1, n2, F(r), P(r), |F(r) - P(r)|
0.000 2 12 0.043 0.000 0.043
10.000 6 12 0.130 0.009 0.121
14.142 8 12 0.174 0.019 0.155
20.000 9 12 0.196 0.037 0.158
22.361 11 12 0.239 0.046 0.193
30.000 13 12 0.283 0.082 0.201
36.056 15 12 0.326 0.116 0.210
41.231 18 12 0.391 0.149 0.242
42.426 21 12 0.457 0.157 0.299
44.721 22 12 0.478 0.173 0.305
50.990 24 12 0.522 0.219 0.303
53.852 28 11 0.596 0.241 0.355
56.569 29 11 0.617 0.262 0.355
58.310 30 11 0.638 0.276 0.362
60.828 31 11 0.660 0.296 0.363
63.246 33 10 0.688 0.316 0.371
80.623 38 8 0.760 0.461 0.299
82.462 39 8 0.780 0.476 0.304
92.195 40 8 0.800 0.554 0.246
94.340 41 8 0.820 0.571 0.249
98.489 43 8 0.860 0.602 0.258
106.301 44 8 0.880 0.658 0.222
107.703 46 6 0.885 0.668 0.217
110.000 47 6 0.904 0.683 0.221
111.803 48 5 0.906 0.695 0.211
114.018 51 3 0.927 0.709 0.218
120.416 52 3 0.945 0.748 0.198
136.015 54 1 0.947 0.827 0.120
162.788 55 0 0.948 0.919 0.029
208.087 57 0 0.983 0.984 0.001
210.238 58 0 1.000 0.985 0.015
The Maximum d(r): 0.371
This d(r) is at 0.010 signficance level
References
Boots, Barry N. and Getis, Arthur, 1988, Point Pattern Analysis, Sage University
Paper series on Quantitative Applications in the Social Sciences, series no.07-001, Beverly Hills: Sage Publications
Diggle, Peter J., 1990, "A Point Process Modelling Approach to Raised Incidence of a Rare Phenomenon in the Vicinity of a Prespecified Point." Journal of the Royal Statistical Society A, 153(3), 349-362